Analysing segmentation from other napari plugins#
Introduction#
While manual analysis of regions can be effective for various applications, there are instances where automated methods are necessary. In such cases, other napari plugins can be used.
Plugins exist to segment many features common to biomedical images, and their results can be incorporated with BrainGlobe to analyse the distribution of features of interest within the context of an anatomical atlas.
To be compatible with brainglobe-segmentation, the third-party napari plugin must be able to take a 3D image layer as
input and return one of:
A 3D labels layer with either a 2 or 3D region labeled (e.g., segmenting bulk axonal projections)
A 3D points layer with a series of points corresponding to a trajectory through 3D space
Hint
It is possible to segment structures outside the brainglobe-segmentation plugin and import them in later (e.g., by
saving to a .tiff and reloading). However, it is simpler to load the data using brainglobe-segmentation and then segment
it using a third-party plugin. This ensures that the coordinate spaces you are using are consistent.
Instructions#
Segmenting your feature of interest#
Open napari
Install the napari plugin you require to segment your data. To find out which plugins are available, see the napari hub.
Use the plugin to segment your feature of interest. Ensure it returns a 3D labels or points layer.
Hint
There are many plugins available, and we can’t recommend a single tool for all data. For simple segmentation of 3D
structures we often use
napari-simpleitk-image-processing. Another
useful tool (also by Robert Haase) is
napari-segment-blobs-and-things-with-membranes.
Analyse the segmented layer#
Hint
At this point you could load a layer segmented previously and saved to disk
If required, rename the layer to be analysed
Click either the
Track tracingorRegion segmentationbuttons to load the respective panels.Highlight the layer to be analysed in the napari layers list:
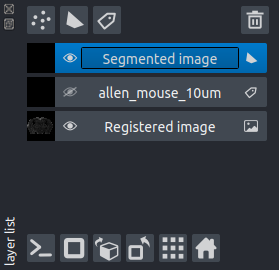
Click either the
Add track from selected layerorAdd region from selected layerbutton as applicable. This will add the region/points from the layer to thebrainglobe-segmentationanalysis list (in the same way as if it had been analysed manually withinbrainglobe-segmentation).Follow the instructions for either:
Note
All data will be saved into your brainreg output directory
